作 者:Shen, YJ; Chalopin, D; Garcia, T; Boswell, M; Boswell,W; Shiryev, SA; Agarwala, R; Volff, JN; Postlethwait, JH; Schartl, M; Minx, P;Warren, WC; Walter, RB 影响因子:3.965
刊物名称:BMC GENOMICS
出版年份:2016
DOI:10.1186/s12864-015-2361-z
Background:Xiphophorusfishes are represented by 26 live-bearing species of tropical fish that express many attributes (e.g., viviparity, genetic and phenotypic variation, ecological adaptation, varied sexual developmental mechanisms, ability to produce fertile interspecies hybrids) that have made attractive research models for over 85 years. Use of various interspecies hybrids to investigate the genetics underlying spontaneous and induced tumorigenesis has resulted in the development and maintenance of pedigreedXiphophoruslines specifically bred for research. The recent availability of theX. maculatusreference genome assembly now provides unprecedented opportunities for novel and exciting comparative research studies amongXiphophorusspecies.
Results: We present sequencing, assembly and annotation of two new genomes representingXiphophorus couchianusandXiphophorus hellerii. The finalX. couchianusandX. helleriiassemblies have total sizes of 708 Mb and 734 Mb and correspond to 98 % and 102 % of theX. maculatusJp 163 A genome size, respectively. The rates of single nucleotide change range from 1 per 52 bp to 1 per 69 bp among the three genomes and the impact of putatively damaging variants are presented. In addition, a survey of transposable elements allowed us to deduce an ancestral TE landscape, uncovered potential active TEs and document a recent burst of TEs during evolution of this genus.
Conclusions: Two newXiphophorusgenomes and their corresponding transcriptomes were efficiently assembled, the former using a novel guided assembly approach. Three assembled genome sequences within this single vertebrate order of new world live-bearing fishes will accelerate our understanding of relationship between environmental adaptation and genome evolution. In addition, these genome resources provide capability to determine allele specific gene regulation among interspecies hybrids produced by crossing any of the three species that are known to produce progeny predisposed to tumor development.
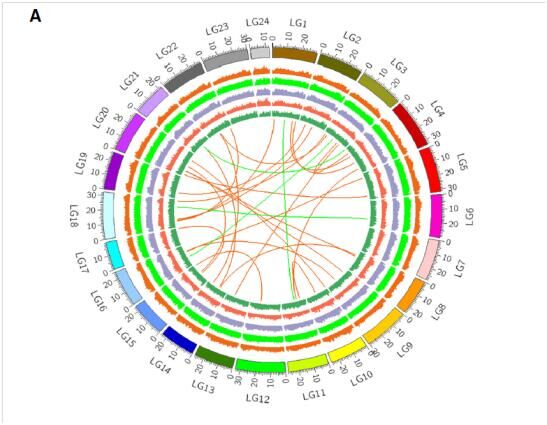
Fig. 2.Distribution of polymorphisms inXiphophorusgenome among 24 chromosomes.aThe histogram rings in the Circos plot represent the number of SNCs in 100 kb bins normalized by 3000. Tracks from outside circles to inner circles are polymorphisms betweenX. maculatusandX. couchianus(orange), betweenX. maculatusandX. hellerii(light green), only inX. maculatus(purple), only inX. couchianus(red) and only inX. hellerii(dark green). The connecting links in the inner circle show the inter-chromosomal rearrangements betweenX. maculatusandX. couchianus(orange links) andX. maculatusandX. hellerii(green links).

