Lin X.L., Hetharua B., Lin L., Xu H., Zheng T.L., He Z.L. and Tian Y.. 2019. Microbial Ecology, 78(1): 57-69.
Microorganisms play important roles in mangrove ecosystems. However, we know little about the ecological implications of mangrove microbiomes for high productivity and the efficient circulation of elements in mangrove ecosystems. Here, we focused on mangrove sediments located at the Yunxiao National Mangrove Reserve in southeast China, uncovering the mangrove microbiome using the 16S rRNA gene and shotgun metagenome sequencing approaches. Physicochemical assays characterized the Yunxiao mangrove sediments as carbon (C)-rich, sulfur (S)-rich, and nitrogen (N)-limited environment. Then phylogenetic analysis profiling a distinctive microbiome with an unexpected high frequency of Chloroflexi and Nitrospirae appeared to be an adaptive characteristic of microbial structure in S-rich habitat. Metagenome sequencing analysis revealed that the metabolic pathways of N and S cycling at the community-level were routed through ammonification and dissimilatory nitrate reduction to ammonium for N conservation in this N-limited habitat, and dissimilatory sulfate reduction along with polysulfide formation for generating bioavailable S resource avoiding the biotoxicity of sulfide in mangrove sediments. In addition, methane metabolism acted as a bridge to connect C cycling to N and S cycling. Further identification of possible biogeochemical linkers suggested Syntrophobacter, Sulfurovum, Nitrospira, and Anaerolinea potentially drive the coupling of C, N, and S cycling. These results highlighting the adaptive routed metabolism flow, a previously undescribed property of mangrove sediment microbiome, appears to be a defining characteristic of this habitat and may significantly contribute to the high productivity of mangrove ecosystems, which could be used as indicators for the health and biodiversity of mangrove ecosystems.
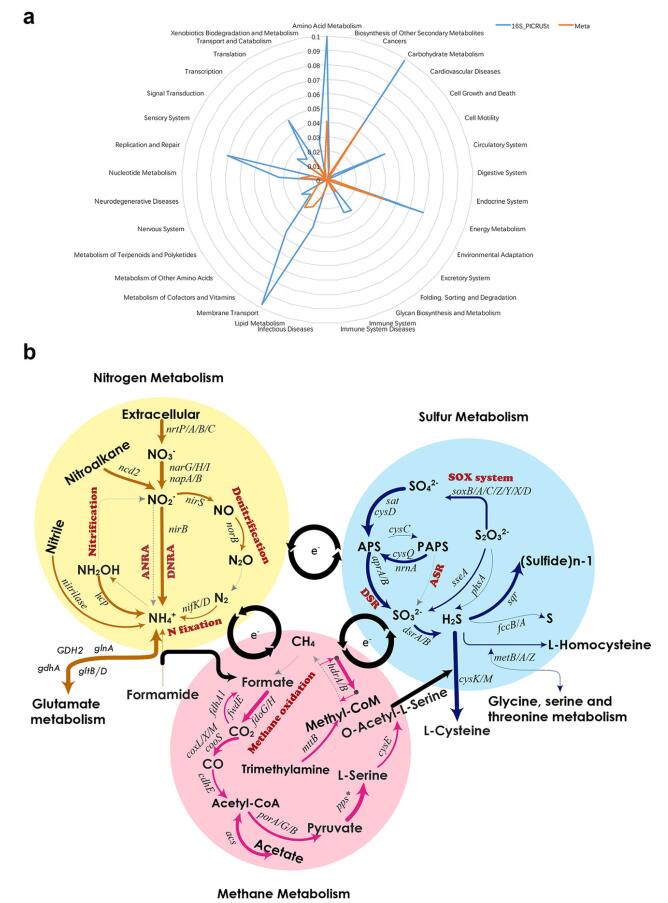
Figure 1. KEGG functional categories (a) and pathways (b) related to methane, nitrogen, and sulfur in Smetagenome. The top 30, 25, and 30 KEGG characteristics identified in methane, N, and S metabolisms and corresponding transformation processes are shown. The gene with an asterisk (pps*) represents a series of genes involving the conversion from pyruvate to Lserine. The width of the arrows indicates the relative abundance of the related genes in the metagenome. The processes labeled with gray dotted lines indicate the presence in the metagenome or predicted datasets but with a low frequency. The black arrows show the putative coupling mechanisms among methane, N, and S cycles: matter flows via formate and O-acetyl-L-serine, and electron transfer via respiration processes.

