Ma D.N., Guo Z.J., Ding Q.S., Zhao Z.Z., Shen Z.J., Wei M.Y., Gao C.H., Zhang L.D., Li H., Zhang S., Li J., Zhu X.Y. and Zheng H.L.. 2021. Molecular Ecology Resources, 21(5):1593-1607.
Aegiceras corniculatum is a major mangrove plant species adapted to waterlogging and saline conditions, grows in the coastal intertidal zone of tropical and subtropical regions. Here, we present a chromosome-level genome assembly of A. corniculatum by incorporating PacBio long-read sequencing and Hi-C technology. The results showed that the PacBio draft genome size is 906.63 Mb. Hi-C scaffolding anchored 885.06 Mb contigs (97.62% of draft assembly) onto 24 pseudochromosomes. The contig N50 and scaffold N50 were 7.1 Mb and 37.74 Mb, respectively. Out of 40,727 protein-coding genes predicted in the study, 89% have functional annotations in public databases. We also showed that of the 603.93 Mb repetitive sequences predicted in the assembled genome, long terminal repeat retrotransposons constitute 41.52%. The genome evolution analysis showed that the A. corniculatum genome experienced two whole-genome duplication events and shared the ancient gamma whole-genome triplication event. A comparative genomic analysis revealed an incidence of expansion in 1,488 gene families associated with essential metabolism and biosynthetic pathways, including photosynthesis, oxidative phosphorylation, phenylalanine, glyoxylate, dicarboxylate metabolism, and DNA replication, which probably constitute adaptation traits that allow the A. corniculatum to survive in the intertidal zone. Also, the systematic characterization of genes associated with flavonoid biosynthesis pathway and the AcNHX gene family conducted in this study will provide insight into the adaptation mechanism of A. corniculatum to intertidal environments. The high-quality genome reported here can provide historical insights into genomic transformations that support the survival of A. corniculatum under harsh intertidal habitats.
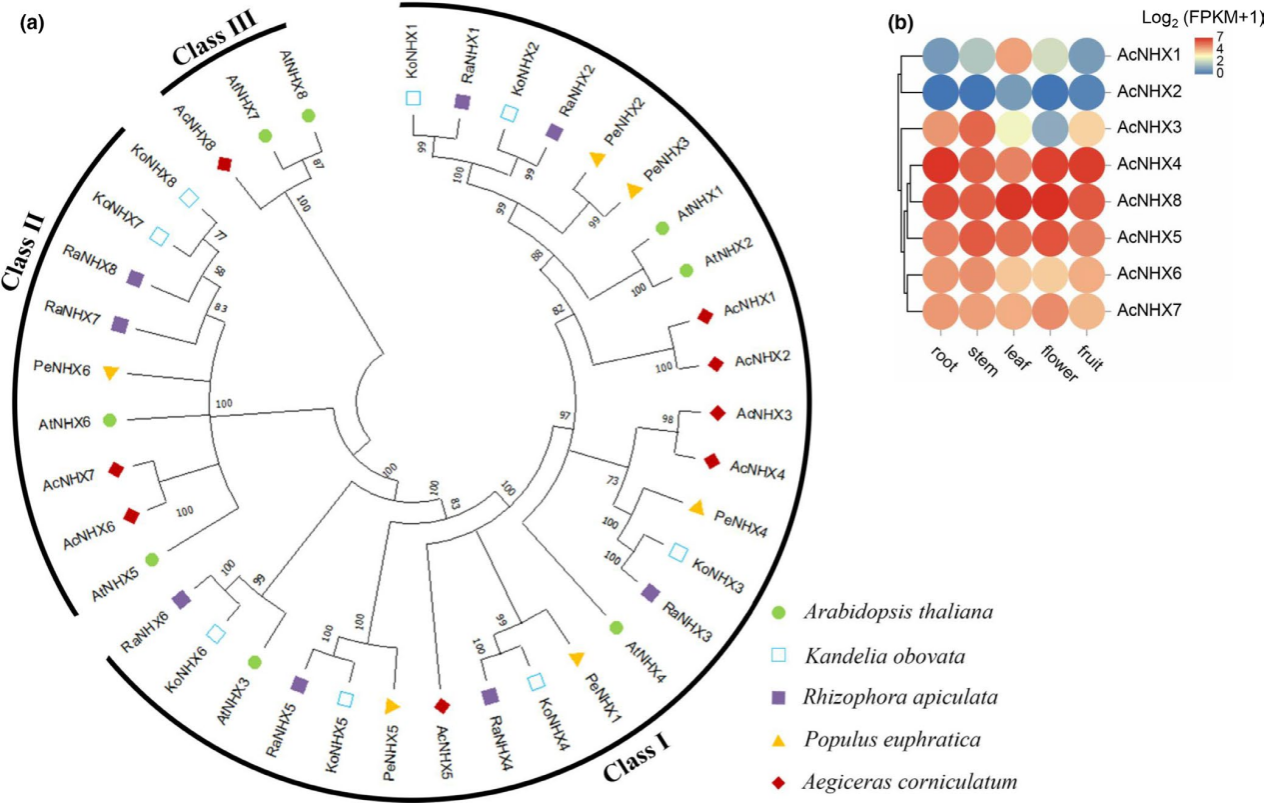
Figure 1. Analysis of NHX gene family. (a) Phylogenetic analysis of NHX genes from Arabidopsis thaliana (At), Kandelia obovata (Ko), Rhizophora apiculata (Ra), Populus euphratica (Pe), and A. corniculatum (Ac). There are three main categories: Class I, Class II and Class III (b) Heatmap showing the expression pattern of AcNHX genes in different tissues including root, stem, leaf, flower and fruit.

